The Missing Information in Addressing Ongoing Measles Outbreaks
Why Identifying the Measles Genotype Matters for Tracing Outbreak Origins
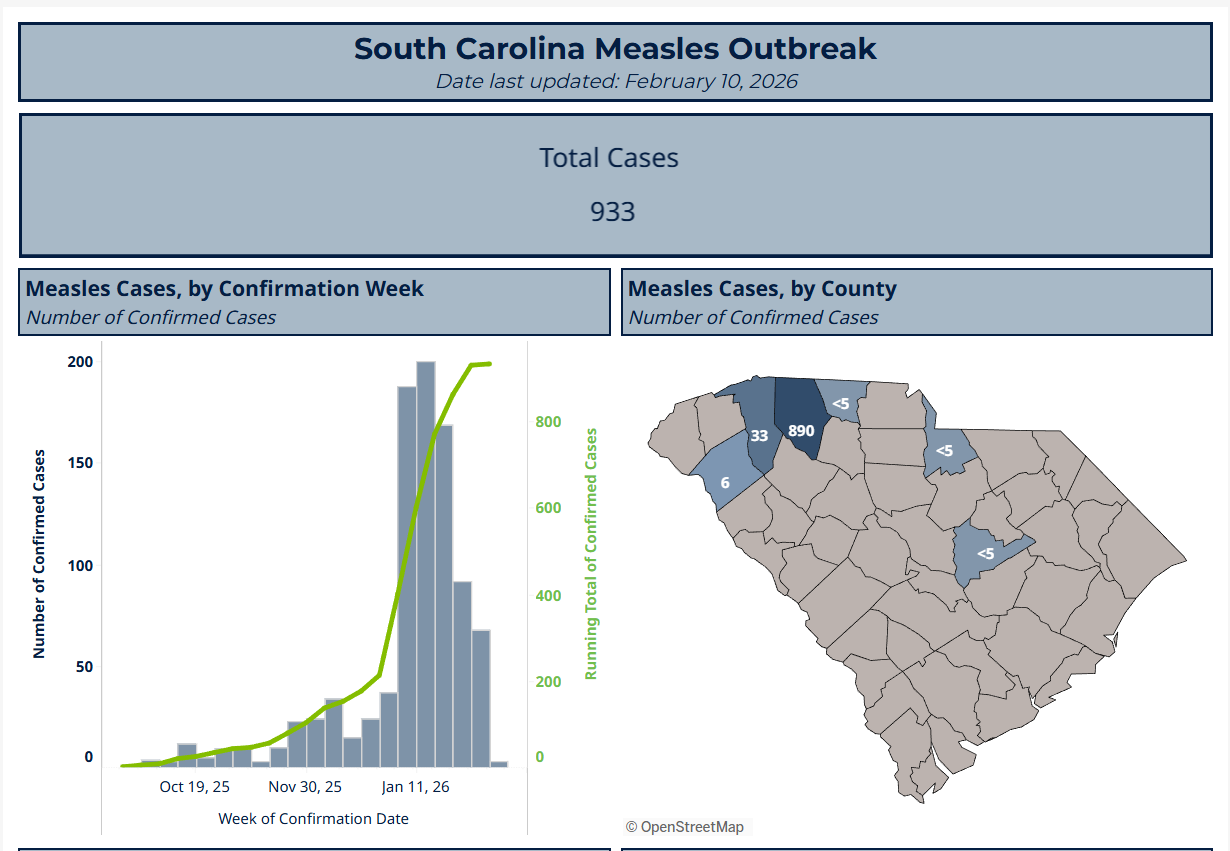
As of early February 10, 2026, South Carolina remains the center of the largest measles outbreak in the US in over 30 years, with 933 confirmed cases. However, there is some key information about this outbreak that is not being reported to the public.
Source: https://dph.sc.gov/diseases-conditions/infectious-diseases/measles-rubeola/measles-dashboard
The health department dashboard shows the epidemiological curve for the outbreak, as well as; measles cases by county, vaccination status and age group.
🌏Knowing the genotype of a measles strain can help one understand where measles outbreaks are coming from
Genotyping was established in 1998 and, with over 50,000 sequence submissions to the Measles Nucleotide Surveillance database, has proven to be an effective resource for countries attempting to trace pathways of transmission.
Epidemiologic investigations are supposed to identify where an outbreak started, how it is spreading, and what can be done to stop it. In the case of measles, that means determining the specific viral genotype involved. Because measles genotypes are linked to particular regions of the world, identifying the strain responsible is critical to tracing the true source of an outbreak. Without that information, the public is left with incomplete answers about where cases originated and how transmission is actually occurring.
🧬Measles is caused by a single serotype of the virus
While the claim is that measles virus is a single serotype, over 24 genotypes, divided into clades A–H, have been identified.
Wild-Type Measles Strains (Genotypes):
These 19 genotypes have been detected since 1990:
A*, B2, B3, C1, C2, D2, D3, D4, D5, D6, D7, D8, D9, D10, D11, G2, G3, H1, H2
These 4 genotypes were identified by global surveillance since 2018:
B3, D4, D8, H1
*Note: All vaccine strains are genotype A
Vaccine Strains:
Edmonston Strain: The foundation for most modern vaccines, including the common Edmonston-Zagreb, Schwarz, and Moraten strains.
Moraten Strain: The primary strain used in the measles-mumps-rubella (MMR) vaccine in the United States.
Other Vaccine Strains: Included are CAM-70, TD 97, and Shanghai 191.
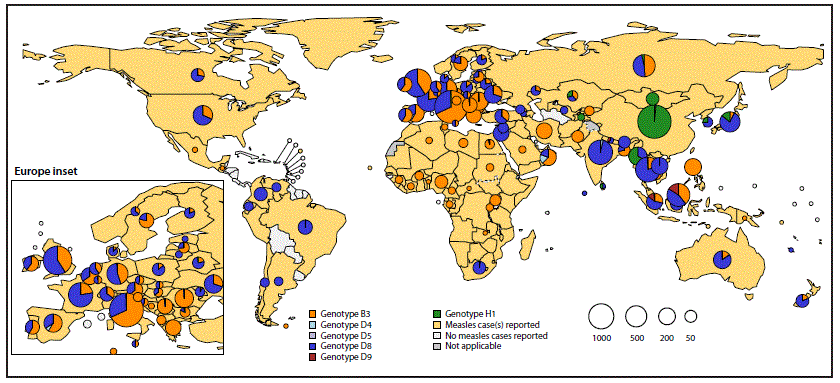
The map shows the global distribution of measles virus genotypes circulating between 2016 and 2018. The size of the circles reflects the numbers of replicates reported for each genotype therefore, the larger circle indicates that a large proportion of measles viruses in that region were that denoted genotype.
Source: https://www.cdc.gov/mmwr/volumes/68/wr/mm6826a3.htm
While vaccine strains are mostly genotype A, wild-type strains like D8, B3, and H1 circulate globally.
Genotype D8: Currently the most commonly detected wild-type strain in many regions, including recent US outbreaks.
Genotype B3: Frequently detected, especially in parts of Africa and the Middle East.
Genotype H1: Common in the Western Pacific region, particularly China.
Source: https://www.mdpi.com/2076-393X/12/7/810#vaccines-12-00810-f002
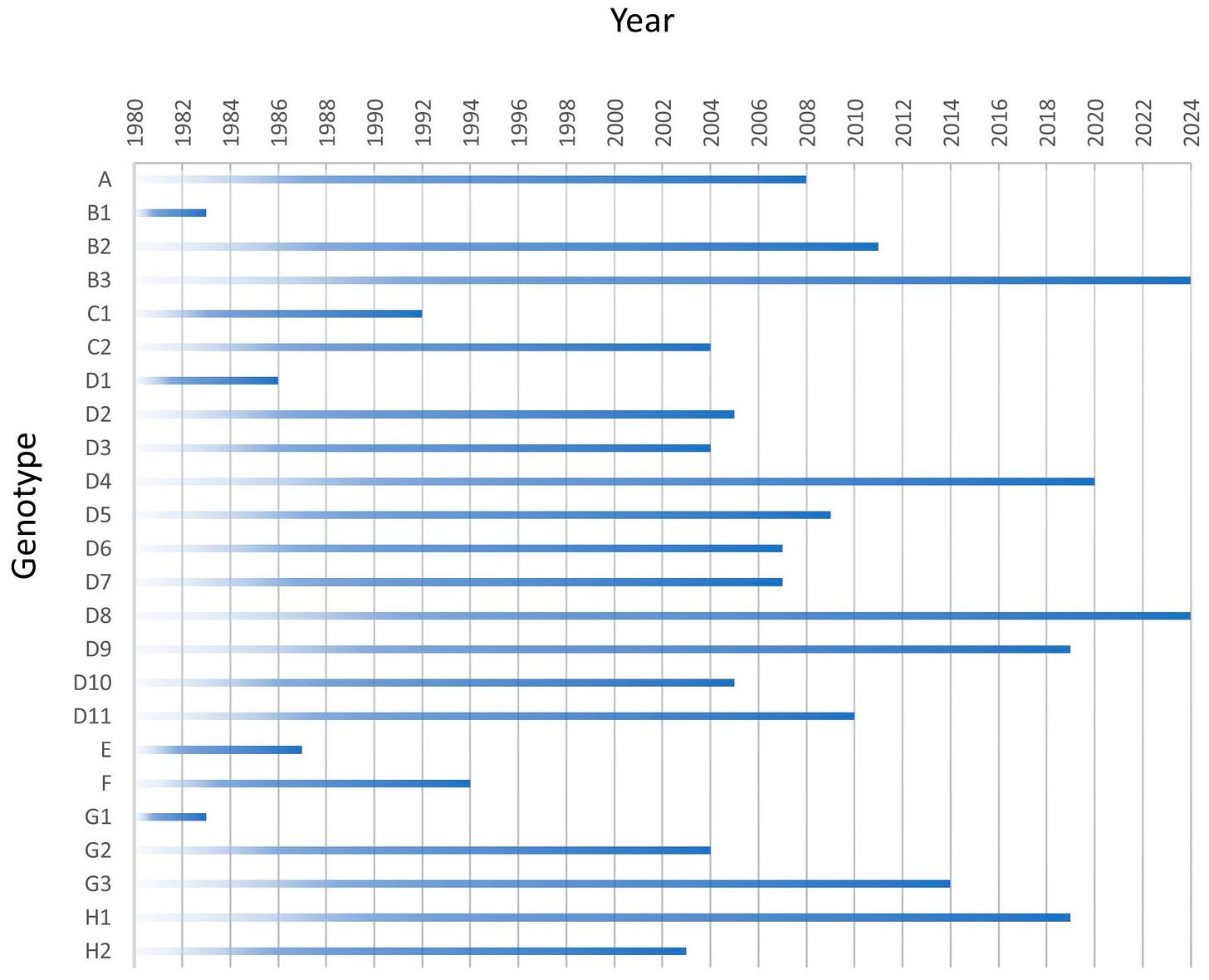
This chart shows the year of last detection of measles genotypes. The last year of documented circulation reported to Measles Nucleotide Surveillance database for all 24 genotypes is shown. Viruses with a date of 2024 are currently circulating. All other viruses have had transmission interrupted in the year depicted.
🦠Outbreaks in the US are imported from other countries
The 2014–2015 Disneyland measles outbreak was a high-profile public health event that began in December 2014. The outbreak infected approximately 147 people in the US. Although the original source (index case) was never identified, health officials believe the virus was brought to the park by an infected international traveler. Testing identified measles genotype B3, the same strain that caused a massive outbreak in the Philippines in 2014.
During a May 2017 measles outbreak in Minnesota, a communication’s director for the Minnesota Department of Health confirmed that “that the virus strain making people sick in this outbreak is the B3 wild-type virus.” As shown on the map above, this strain commonly circulates in Africa. US-born children of Somali descent (Somali children) accounted for 55 (85%) of the cases.
More recently, ten measles outbreaks have been reported in 2025; 751 (94%) of all reported confirmed measles cases were outbreak-associated. An imported source was identified for seven outbreaks, and the source of three outbreaks remains unknown. Forty-eight (6%) cases were directly imported from other countries. Source countries of the 48 imported measles cases included Canada (10 cases), Vietnam (10), Mexico (seven), Pakistan (three), the Philippines (two), Saudi Arabia (two), and one imported case each from Afghanistan, Australia, Guinea, Netherlands, Somalia, Spain, and Uganda; a source country could not be determined for seven travelers who visited multiple countries during their exposure period.
The largest outbreak began among a close-knit community Gaines County, Texas in January 2025 and has accounted for 654 (82%) cases reported during 2025; the source of this outbreak remains unknown. The CDC reported 128 measles genetic sequences. Texas submitted 92 identical sequences that belong to the D8 genotype. Ten identical sequences have been reported from New Mexico, and one sequence matching the Texas outbreak virus was reported from Kansas. Five distinct B3 genotype sequences have been reported from eight other states.
The Arizona-Utah border region, particularly in the northern region/Mojave County, has experienced a surge in cases, with over 120 people infected by late October 2025. With over 200 cases reported, the D8-9171 strain, in particular, has been identified as the dominant strain.
Outbreak Context: These outbreaks are often started by international travelers and spread rapidly in communities with lower vaccination coverage.
💉Genotyping can also distinguish whether a person has wild-type measles virus infection, or a rash caused by a recent measles vaccination.
During the measles outbreak in California in 2015, a large number of suspected cases occurred in recent vaccinees. Of the 194 measles virus sequences obtained in the United States in 2015, 73 were identified as vaccine sequences (R. J. McNall, unpublished data).
The CDC acknowledges that 5–7% of individuals vaccinated with the live-attenuated measles virus develop a rash and fever post-vaccination, they can meet the clinical case definition for measles if genotype testing is not performed. In addition, in a previous post I outlined how shedding of vaccine virus can occur for up to 29 days after vaccination with documented transmission in households, siblings, sexual partners, and through breastfeeding.
The CDC recommends children receive two doses of the MMR vaccine: the first at 12–15 months and the second at 4–6 years.
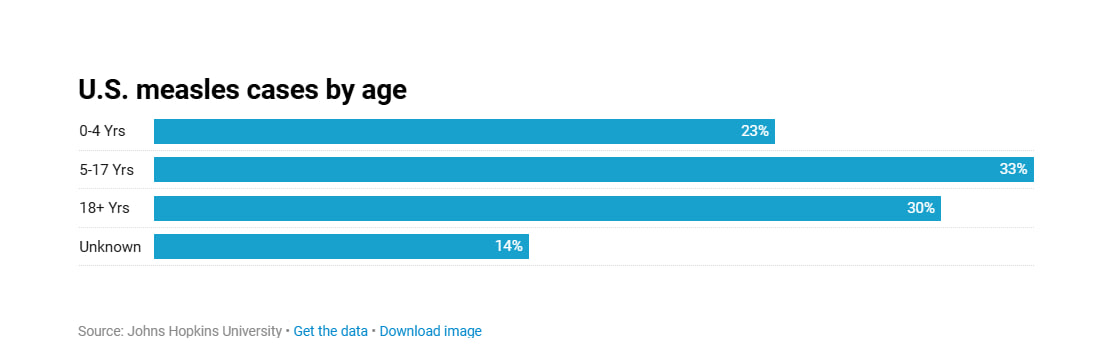
A majority of cases in the South Carolina outbreak are among the 5-11 age group while overall cases in the US have been among the 5-17 age group.
It is interesting to note that back-to-school vaccine clinics in South Carolina are generally held annually in late July through August, leading up to the start of the school year, with additional opportunities often available in early September. In July, South Carolina had two confirmed cases of measles. It was reported the first measles case was associated with exposure while traveling. The second case was a known close contact of the first case. “Earlier in September, South Carolina had one confirmed case of measles also associated with exposure while traveling and was unrelated to the July cases. The current outbreak in the Upstate involves two cases with no history of travel or identified source of exposure.” as stated in a press release on October 2, 2025.
In January 2026 alone, over 1,200 infants aged 6–11 months were vaccinated in the South Carolina outbreak region. If any developed rash within 21 days and were not tested using genotype assays or MeVA RT-qPCR (which detects vaccine-strain genotype A), they may have been misclassified as measles cases.
**Information recently obtained by a request to the South Carolina Department of Public Health revealed that genotype D8 was identified for the majority of positive specimens collected from the outbreak.
💉Does the measles vaccine really cover all genotypes of measles?
The general consensus seems to be that the measles vaccine works against all genotypes of measles because there is only one serotype of measles virus. When a virus is of the same serotype, they have the same antigens—proteins that our bodies recognize and make an immune response to—on the outside.
Measles viruses are characterized based on the sequences of their hemagglutinin (H) and nucleocapsid (N) gene-coding sequences. The primary antigenic surface proteins of the measles virus (MeV) are the H protein and the Fusion (F) protein. The H protein, responsible for receptor binding (to CD46, SLAMF1, or nectin-4), is the main target for neutralizing antibodies.
However, neutralization studies using vaccine-induced H-specific monoclonal antibodies (mAbs) have shown that most of these mAbs distinguish between MV-H variants whereas only a few mAbs neutralize all variants. This would suggest that the basic assumption that measles virus is one serotype is incorrect.
Another study revealed that strains carrying the classical hemagglutinin type predominated until the early 1980s when it became completely replaced with strains possessing the contemporary hemagglutinin type. Researchers identified differences in key antigenic sites on the hemagglutinin protein—areas of the virus that antibodies recognize and bind to—demonstrating that antigenic variation can occur in measles viruses over time.
In plain terms, this means antibodies generated against one genotype may not bind as effectively—or in some cases, as strongly—to other genotypes.
Because neutralizing antibodies are directed primarily against conformational epitopes on the H protein, substitutions within or adjacent to recognized antigenic sites have the potential to alter antibody binding affinity. In principle, antibodies elicited by exposure to one genotype could exhibit differential neutralization efficiency against heterologous genotypes if antigenic determinants are sufficiently altered.
The currently licensed measles vaccines are derived from genotype A strains. Protection across genotypes has historically been attributed to the relative conservation of neutralizing epitopes on the H protein and the functional constraints limiting extensive antigenic drift. Nonetheless, continuous molecular surveillance, antigenic characterization, and neutralization studies remain important to confirm that vaccine-induced sera maintain broad cross-genotypic neutralizing capacity against circulating strains.
Conclusions:
Measles genotyping is not a peripheral laboratory exercise—it is a central component of outbreak investigation and control. Because measles virus genotypes are geographically structured and genetically distinct, molecular characterization helps distinguish importations from sustained transmission chains and provides clarity about outbreak origin. Without genotypic data, assessments of transmission dynamics remain incomplete.
Although measles virus is considered antigenically monotypic and current genotype A–derived vaccines have historically demonstrated broad cross-neutralization of circulating genotypes, documented amino acid variation in key antigenic regions of the hemagglutinin protein underscores the importance of continued molecular and antigenic surveillance. Genotyping alone does not determine vaccine effectiveness, but it provides essential epidemiologic context for interpreting outbreak patterns and immune protection.
Measles outbreaks in the past have been associated with importation of the virus from international visitors but also with fully vaccinated individuals. In an outbreak in New York City in 2011 not only did a fully vaccinated person contract measles, but also spread it to others who themselves were also fully vaccinated. While breakthrough cases can occur for several reasons, including primary vaccine failure (failure to mount adequate immunity after vaccination), secondary vaccine failure (waning antibody levels over time), unusually high exposure intensity, or host-specific immune variability; demonstrating diminished cross-protection due to antigenic divergence would require systematic neutralization studies comparing vaccine-induced sera against contemporary circulating genotypes.
Ongoing surveillance also serves a forward-looking purpose. Should accumulating genetic or antigenic data ever demonstrate meaningful reductions in cross-neutralization against circulating strains, such findings would inform scientific evaluation of whether updating the vaccine seed strain is warranted. Sustained transparency in sequence reporting, neutralization assays, and global strain monitoring remains critical to ensuring that vaccine-induced immunity remains aligned with viruses in circulation.